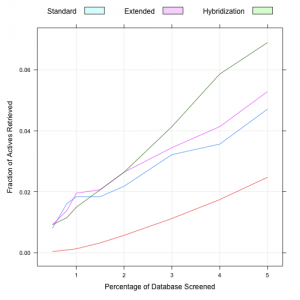
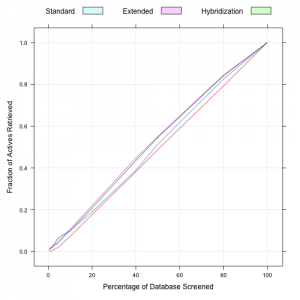
This morning Egon reported that he had implemented a new fingerprinter for the CDK, which only considered hybridization rather than looking at aromaticity. As a result this approach does not require aromaticity perception. I took a quick look to see how it performs in a virtual screening benchmark. Firstly, it’s faster than the other CDK hashed fingerprints – 15,030 fingerprint calculations took ~ 60s with the hybridization only fingerprint. In contrast the extended fingerprint took 80s for the same set of molecules. To test the utility of the fingerprint in a virtual screening scenario I evaluated enrichment curves (see here for a comprehensive comparison of CDK fingerprints) using the AID 692 MUV benchmark dataset. The plots below show the enrichment curves for the first 5% of the database and the entire database. The red curve corresponds to random selections. (In this experiment the database consists of 15,000 decoys and 30 actives). The enrichment factor for the standard, extended and hybiridization only fingerprints were 0.94, 1.06 and 1.38 respectively.
Overall, the hybridization only fingerprint performs comparably to the extended fingerprint and better than the standard one. But at a small percentage of the database screened, it appears that this fingerprint outperforms both. Of course, this is only one dataset, and more MUV datasets should be analyzed to get a more comprehensive view.