In a previous post I described how one requires a custom RecordReader class to deal with multi-line records (such as SD files) in a Hadoop program. While it worked fine on a small input file (less than 5MB) I had not addressed the issue of “chunking” and that caused it to fail when dealing with larger files (the code in that post is updated now).
When a Hadoop program is run on an input file, the framework will send chunks of the input file to individual RecordReader instances. Note that it doesn’t actually read the entire file and send around portions of it – that would not scale to petabyte files! Rather, it determines the size of the file and ends start and end offsets into the original file, to the RecordReaders. They then seek to the appropriate position in the original file and then do their work.
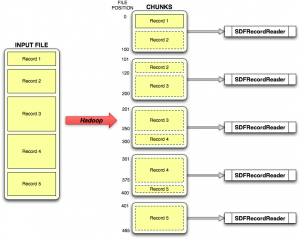
The problem with this is that when a RecordReader receives a chunk (defined in terms of start and offsets), it can start in the middle of a record and end in the middle of another record. This shown schematically in the figure, where the input file with 5 multi-line, variable length records is divided into 5 chunks. As you can see, in the general case, chunks don’t start or end on record boundaries.
My initial code, when faced with chunks failed badly since rather than recognizing chunk boundaries it simply read each record in the whole file. Alternatively (and naively) if one simply reads up to a chunk boundary, the last and first records read from that chunk will generally be invalid.
The correct (and simple) strategy for an arbitrary chunk, is to make sure that the start position is not 0. If so, we read the bytes from the start position until we reach the first end of record marker. In general, the record we just read will be incomplete, so we discard it. We then carry on reading complete records as usual. But if, after reading a record, we note that the current file position is beyond the end position of the current chunk, we note that the chunk is done with and just return this last record. Thus, according to the figure, when processing he second chunk from the top, we read in bytes 101 to 120 and discard that data. We then start reading the initial portion of Record 3 until the end of the record, at position 250 – even though we’ve gone beyond the chunk boundary at position 200. However we now flag that we’re done with the chunk and carry on.
When another RecordReader class gets the next chunk starting at position 200, it will be dumped into the middle of Record 3. But, according to our strategy, we simply read till the end of record marker at position 250 and discard the data. This is OK, since the RecordReader instance that handled the previous chunk already read the whole of Record 3.
The two edge cases here are when the chunk starts at position 0 (beginning of the input file) and the chunk ends at the end of file. In the former case, we don’t discard anything, but simply process the entire chunk plus a bit beyond it to get the entire last record for this chunk. For the latter case, we simply check whether we’re at the end of the file and flag it to the nextKeyValue() method.
The implementation of this strategy is shown in the SDFRecordReader class listing.
In hindsight this is pretty obvious, but I was bashing myself for a while and hopefully this explanation saves others some effort.
Just a short question on the special case when a chunk happens to end with the \$$$$
\ string.
I couldn’t really understand from the code how that would be handled, more specific if fsin.getPos() < end in that case.
I suppose that otherwise there is a chance of loosing the following record.
/Fredrik
To understand this you need to consider how key/values are called – it’s a lot like implementing Iterator in Java.
Hadoop will call nextKeyValue() to check whether there is a new key/val pair and if so it will call getCurrentKey() and getCurrentValue().
So when we hit the $$$$ at the end of the file, readUntilMatch returns false, since fsin.getPos = end. Back in nextKeyVal() this will cause stillInChunk to be set to false and the method returns true – indicating that there is a key/value pair available (the last one in the file).
When nextKeyValue() is called again, the test
if (!stillInChunk) return false
will cause the method to return false and hence the caller will understand there are no more key/val pairs available
Thanks for the great post, it helped a lot. Just a quick question:
If we try to read past our chunk boundary, won’t we face the risk of having to read from a remote machine, therefore violate data locality and face performance issues?
Ahmet, yes, I think you’re right. I’m not sure how one would get around this.
Thanks for the post.It really helped to understand how hadoop ensures discarding of partial read of records.
However I am wondering what will happen when $$$$ is the part of my actual data?
In that case the method breaks down. For SDF files, $$$$ is defined as the record separator, so I wouldn’t anticipate this problem with that type of file
I too wondering how it all continue to read from next split as in start of the code we have initialized the FSDataInputStream object with reference to incoming split path [which eventually will not be the path for next split]
Path path = split.getPath();
FileSystem fs = path.getFileSystem(conf);
fsin = fs.open(path);